WIP: Towards Better Research Management Tools
As part of my day job or my personal research I need to find answers to scientific/engineering questions such as:
- “what methods exist to extract information from structure”,
- “what are the effects of parametric knowledge on the factuality of summaries”
- “what’s a good definition for domain and how does that correlate to performance”
Finding research papers to answer a given research question can be a pain in my experience. This is partly due to the headaches of needing many tools to find all relevant results and the pain of collating/organizing those results.
IMO there are four steps to finding papers:
- Problem Definition: Defining the scope (i.e., what papers/fields/subjects/topics are relevant to my research question, survey)
- Selection: Finding (all) relevant papers within that scope
- Curation: Triaging, collating, and organizing relevant papers
- Interpretation: Interpreting results; either finding a conclusion, refining scope (i.e., rinse, wash, and repeat), or identifying further research needed.
While each step poses challenges, 2 and 3 are unnecessarily hard given the current technology.
The Status Quo
Currently searching for papers is frustrating because I need to use different tools (i.e., selection) and organize the results manually (i.e., curation). I’ve tried different combinations of search methods, namely
- Google (search and its other products)
- arXiv
- Semantic Scholar (closest to how I like to organize results)
- alphaXiv (as an aside I like their UX, especially how they integrate multiple search methods)
- Connected papers
- Deep research tools (e.g., deep research)
Aaron Tay gives some insight into how the opaqueness of current search approaches is partly what makes finding relevant papers harder. In short, under the hood each of these tools uses different technologies such as keyword search (e.g., boolean search, BM25), neural embeddings, citation-based methods, and now agents. Each method expects different input formats and is lacking in slightly different ways. In my experience that translates into needing a large number of tools to perform an exhaustive search to answer a given question or perform a survey. Due to the differences in capabilities and the strengths of each not being well communicated, it can be difficult to know when a poor result is due to the query, the method, or the tool itself.
Here’s a list of the failure modes of these different approaches in my experience.
- Keyword based methods are pretty intuitive and give great results… as long as you have the correct keywords…
- Neural methods can be used to perform QA, IR, amongst other tasks. In general they have a fair amount of flexibility when it comes to finding results, i.e., not just keyword based. That said they are often trained to perform a specific task on specific data and are likely to fail in unexpected ways outside of their domain. My favorite failure mode so far has been ranking punctuation heavy spans very highly.
- Bibliographic methods are cool and leverage the work done by scientific communities through citation tracking. That said they tend to not be interdisciplinary and can wind up only finding certain pools of authors that co-cite each other’s work. If there are cliques, veins of research, or different communities you may not find them unlike textual methods. Moreover once in a while papers irrelevant to your query can be surfaced if they are just oft-cited.
- Deep research/agentic approaches are relatively new. In general I’ve been disappointed by these tools,
they tend to pull in only tangentially relevant results and miss some clear matches. I’m not quite sure
how they work or if they perform the work consistently because
- tool calls aren’t always shown
- reasoning traces can be misleading
- generation is random and non-deterministic, even under temperature=0
Each method requires an understanding of their potential shortcomings, which in practice means all these tools need to be used together. Taking a step back, this means different interfaces, tools, and then needing to manually combine results. In other words, it’s time to choose a system to organize your search results :) Some examples include
- An Excel sheet (I still use this, it’s flexible, simple, shareable, and pretty universal)
- Zettelkasten
- Zotero
- Semantic Scholar (I like the idea of this tool more than the tool itself)
I’ve tried different systems above and the only one I’ve consistently used is Excel. I hate it, copying in results and correcting the occasional gaff gets tedious.
Custom Solution
In short there are two problems I want to tackle; selection and curation. The selection subsection will discuss primarily the back-end and search methods made available in the tool. The curation section will cover the UI/UX around collating data and recommendation engines used to help find additional results.
Selection
I think it’s easy to fall into the trap of a better search method being all that’s needed, but I don’t think that’s the case here. Rather than try to create a perfect search method, the approach I chose is to offer more search options and methods in one spot (i.e., site) and see what works vs what doesn’t in the long term. To be clear, I don’t think this search experience is for everyone. There are a lot of choices that can be made and honestly they will probably make the process more confusing (even for me having more than 2-3 choices is a lot).
This translates into building a “search engine for papers” (the nth one) and the components required to maintain a search engine. At a high level, there are three parts to making a search tool:
- Corpus ingestion & curation
- Indexing
- User experience
Corpus Ingestion & Curation
Ingestion and curation sounds misleadingly simple. You just find the results and take them down. As I went along there were always additional steps. Just taking a look at the ingestion pipeline below will give you an idea of the scale of work required. As I’m discovering this list is still incomplete, but works well enough for now.
Update Script
# Crawling
kaggle datasets download Cornell-University/arXiv --unzip --force
# Data intake and preprocessing
python insert_raw_data.py
python score_relevance.py
python download_pdfs.py
python extract_pdfs.py
# Data transfer
python insert_docs.py
# Add metadata only to search data
python add_semantic_scholar_metadata.py -t SEARCH_CORPUS_TABLE_DATA
# Span creation
python create_spans.py -t SEARCH_CORPUS_TABLE_DATA -f text -s regex -n 512 -o 0
python create_spans.py -t SEARCH_CORPUS_TABLE_DATA -f text -s regex -n 512 -o 256
python create_spans.py -t SEARCH_CORPUS_TABLE_DATA -f text -s regex -n 256 -o 0
python create_spans.py -t SEARCH_CORPUS_TABLE_DATA -f text -s regex -n 256 -o 128
# Indexing
python embed_db.py -t SPANS_SEARCH_CORPUS_TEXT_REGEX_256_OFFSET_0_DATA -f span_text -m "text-embedding-nomic-embed-text-v1.5"
python embed_db.py -t SPANS_SEARCH_CORPUS_TEXT_REGEX_256_OFFSET_0_DATA -f span_text -m "BM-25" --force
python embed_db.py -t SPANS_SEARCH_CORPUS_TEXT_REGEX_256_OFFSET_128_DATA -f span_text -m "text-embedding-nomic-embed-text-v1.5"
python embed_db.py -t SPANS_SEARCH_CORPUS_TEXT_REGEX_256_OFFSET_128_DATA -f span_text -m "BM-25" --force
python embed_db.py -t SPANS_SEARCH_CORPUS_TEXT_REGEX_512_OFFSET_0_DATA -f span_text -m "text-embedding-nomic-embed-text-v1.5"
python embed_db.py -t SPANS_SEARCH_CORPUS_TEXT_REGEX_512_OFFSET_0_DATA -f span_text -m "BM-25" --force
python embed_db.py -t SPANS_SEARCH_CORPUS_TEXT_REGEX_512_OFFSET_256_DATA -f span_text -m "text-embedding-nomic-embed-text-v1.5"
python embed_db.py -t SPANS_SEARCH_CORPUS_TEXT_REGEX_512_OFFSET_256_DATA -f span_text -m "BM-25" --force
python embed_db.py -t SEARCH_CORPUS_TABLE_DATA -f text abstract -m "BM-25" --force
python embed_db.py -t SEARCH_CORPUS_TABLE_DATA -f abstract -m "text-embedding-nomic-embed-text-v1.5"
python embed_db.py -t SEARCH_CORPUS_TABLE_DATA -f abstract -m "naver/splade-cocondenser-ensembledistil"
# Create graph for bibliometric recommender
python create_citation_graph.py -t SEARCH_CORPUS_TABLE_DATA --force
# Display results
python analyze_pipeline.py
To make sure the dataset stays up to date the code is designed to store state in the database and only process items that have failed or new items. This means that after the first run, the number of documents processed each run is minimal (only new documents and previously failed documents). I use cron to run the update job weekly (as this is the frequency with which the arXiv dataset gets updated).
Cron Job for Running Update Pipeline
0 2 * * 0 source ~/arXiv/arXiv_env/bin/activate && /home/felix/research_assistant/src/intake_pipeline/update_pipeline.sh
Crawling
For a search engine to work, you need papers to search and to keep that corpus up to date. That’s what crawling does. You scour the web for content and download it. I decided to keep it simple and use papers off of arXiv only.
Yes, I’m aware I could use Semantic Scholar. My reasoning behind that choice is:
- I’ve been had by abandoned AI2 tools before (RIP AllenNLP).
- this provides the flexibility to build out my own pipeline in the future
- while publishing preprints isn’t required, it is a common practice so coverage of papers is pretty good IMO. Some well-known papers were just released as preprints
- it’s simple to crawl arXiv and within ToS (if done correctly)
Ok so with the corpus chosen, downloading arXiv preprints is pretty straightforward.
- Download a list of arXiv papers from Kaggle
- Run over the entire list of papers and import all previously unseen (i.e., new) arXiv IDs and versions into a database, specifically the “raw storage table”.
Storage
For storage I opted for a single database to warehouse both raw data and processed data. Honestly there are too many products for vector search; my criteria to whittle down the list was the following:
- SQL database since I wanted to store app data in a relational way as well
- Support multiple indices and types of indices, namely dense vectors, sparse vectors, BM25
- Straightforward to use (i.e., not use multiple programs and need to sync them up)
- Open source
Based on these criteria paradedb (or pgvector) was a clear winner for me.
I had read some complaints about performance, but this may well be old news. Performance seems comparable to Elasticsearch (in certain respects according to this benchmark) and it has been working well. My only complaint is that queries can take 15 seconds+ sometimes, but this may well be the hardware I’m running the DB on (a 2 core dell wyse 5070) and the fact I haven’t indexed the vector columns over 2M+ records.
The architecture of the DB is pretty straightforward, here’s a UML diagram showcasing it: This likely will change with time. For now I wouldn’t over-index on it, just sharing it for those who are interested in it. The highlights are
- A raw preprocessing table where results are warehoused upon intake
- A search corpus
- Span tables that correspond to different span sizes and types
- Metadata tables that track query usage
- A graph citation table that is used to store bibliometric data
UML Relational Diagram
erDiagram
%% Generated from db_schemas.py plus live span-table discovery
%% Missing live foreign keys are kept when pipeline or backend code relies on them
%% QUERY_USAGE_MAP._span_id is polymorphic and is resolved through QUERY_USAGE_MAP._span_table
%% Current backend search indices write _span_table as search_corpus or spans_search_corpus_text_regex_512
%% bibliometry_table is populated from search_corpus in update_pipeline.sh, but its row keys currently reference raw_data.arxiv_id
%% SPANS_TEXT_REGEX_256_OFFSET_0 = spans_search_corpus_text_regex_256_offset_0
%% SPANS_TEXT_REGEX_256_OFFSET_128 = spans_search_corpus_text_regex_256_offset_128
%% SPANS_TEXT_REGEX_512 = spans_search_corpus_text_regex_512
%% SPANS_TEXT_REGEX_512_OFFSET_0 = spans_search_corpus_text_regex_512_offset_0
%% SPANS_TEXT_REGEX_512_OFFSET_256 = spans_search_corpus_text_regex_512_offset_256
%% SPANS_TEXT_WHITESPACE_512 = spans_search_corpus_text_whitespace_512
%% QUERY_USAGE_MAP = _m2m_query_arxiv_paper
RAW_DATA {
text arxiv_id PK
text title
text abstract
text text
text summary
boolean relevance
text _pdf_path
jsonb citation_list
jsonb reference_list
}
SEARCH_CORPUS {
text arxiv_id PK
text title
text abstract
text text
text _pdf_path
integer citation_count
jsonb citation_list
jsonb reference_list
}
SPANS_TEXT_REGEX_256_OFFSET_0 {
text span_id PK
text arxiv_id FK
text span_text
integer span_start
integer span_end
}
SPANS_TEXT_REGEX_256_OFFSET_128 {
text span_id PK
text arxiv_id FK
text span_text
integer span_start
integer span_end
}
SPANS_TEXT_REGEX_512 {
text span_id PK
text arxiv_id FK
text span_text
integer span_start
integer span_end
}
SPANS_TEXT_REGEX_512_OFFSET_0 {
text span_id PK
text arxiv_id FK
text span_text
integer span_start
integer span_end
}
SPANS_TEXT_REGEX_512_OFFSET_256 {
text span_id PK
text arxiv_id FK
text span_text
integer span_start
integer span_end
}
SPANS_TEXT_WHITESPACE_512 {
text span_id PK
text arxiv_id FK
text span_text
integer span_start
integer span_end
}
QUERY_TABLE {
text query_id PK
text query_text
date query_date
text _query_model
}
QUERY_USAGE_MAP {
text mapping_id PK
text query_id FK
text arxiv_id FK
text _span_id
text _span_table
integer rank_position
text feedback
}
BIBLIOMETRY_TABLE {
text pair_id PK
text p1_arxiv_id FK
text p2_arxiv_id FK
integer co_citation_count
integer bc_count
float fused_score
}
RAW_DATA ||--o| SEARCH_CORPUS : "same arxiv_id; relevant subset copied downstream"
QUERY_TABLE ||--o{ QUERY_USAGE_MAP : "query_id"
SEARCH_CORPUS ||--o{ QUERY_USAGE_MAP : "arxiv_id; logical FK used for query logging"
RAW_DATA ||--o{ BIBLIOMETRY_TABLE : "p1_arxiv_id"
RAW_DATA ||--o{ BIBLIOMETRY_TABLE : "p2_arxiv_id"
SEARCH_CORPUS ||--o{ SPANS_TEXT_REGEX_256_OFFSET_0 : "arxiv_id; text regex 256 offset 0"
SEARCH_CORPUS ||--o{ SPANS_TEXT_REGEX_256_OFFSET_128 : "arxiv_id; text regex 256 offset 128"
SEARCH_CORPUS ||--o{ SPANS_TEXT_REGEX_512 : "arxiv_id; text regex 512"
SEARCH_CORPUS ||--o{ SPANS_TEXT_REGEX_512_OFFSET_0 : "arxiv_id; text regex 512 offset 0"
SEARCH_CORPUS ||--o{ SPANS_TEXT_REGEX_512_OFFSET_256 : "arxiv_id; text regex 512 offset 256"
SEARCH_CORPUS ||--o{ SPANS_TEXT_WHITESPACE_512 : "arxiv_id; text whitespace 512"
SPANS_TEXT_REGEX_512 ||--o{ QUERY_USAGE_MAP : "_span_id when _span_table = spans_search_corpus_text_regex_512"
Filtering
Once the raw data has been stored, we still need to choose which “relevant” papers to download the corresponding PDFs for further processing. Some optimistic (i.e., naive) napkin math based on a sample led me to believe I could store all of arXiv. After downloading a hundred thousand PDFs, I realized that was not the case. A heuristic for choosing which arXiv PDFs to download was in turn used:
Relevance Scoring Mechanism
relevant_categories = set(["cs.CL", "cs.FL", "cs.HC", "cs.IR", "cs.LG"])
def score_relevance_v1(items):
relevance_items = [
{
"arXiv_id": item["arXiv_id"],
"relevance": all(
[cat in relevant_categories for cat in item["categories"]]
),
"_relevance_score": float(
all([cat in relevant_categories for cat in item["categories"]])
),
"_relevance_confusion": 0.0,
"_relevance_model": "heuristic_v2",
}
for item in items
]
return relevance_items
After filtering the corpus for NLP-related tags there are ~100K documents. There are still irrelevant documents IMO and some results are missed,
but for now the size required (~350GB for the PDFs and ~150GB for the DB) is manageable.
Downloading and Preprocessing
Once we’ve identified relevant papers, the next step is to download the PDFs. After downloading all relevant PDFs, their text is extracted using PyMuPDF. Papers that have text extracted are migrated to the search_corpus table. Afterwards additional metadata for citations and references is added. This is pulled from Semantic Scholar’s graph API. Building out my citation tree seemed like a bit much to start with, but that is a future project.
While the entire paper’s text can be used for certain search methods, it’s not optimal for all methods. Neural models are trained on certain sized chunks or otherwise see diminishing returns when working with long text. For this reason text needs to be chunked. For now spans of a fixed word length were used (a word as defined by a regex that finds boundaries). These lengths were 128, 256, and 512 words. Spans can be made to overlap using an index. Originally I wanted all spans to be in one table, however I noticed that query speed is directly related to the number of items in a table. Even with an index, tables with a larger number of items are rather slow. So for now every span, offset combination is in a separate table.
Indexing
Ok, so the data has been processed and downloaded. Next step is to make it searchable, this involves indexing the different text fields we want to search.
Earlier I buried the lead a bit. There aren’t just 5 methods of searching, there are hundreds. Beyond the high-level categories I outlined above, each broad category itself contains methods and other variations. Methods can be applied to different fields such as the title, abstract, body, spans, etc. There is also a question of how and when they are applied. Below is a table of combinations currently supported
| End Use Case | Method Family | Technique / Model | Field(s) Indexed | Representation Type | Scoring Type |
|---|---|---|---|---|---|
| Search | Keyword | BM25 | title, abstract, body, spans | Sparse (token-based) | Explicit (BM25: TF-IDF variant) |
| Search | Vector | SPLADE | abstract, spans | Sparse (learned expansion) | Explicit (dot product) |
| Search | Vector | Nomic-1.5 | abstract, spans | Dense | Explicit (cosine / dot product) |
| Search | Exact | SQL equality / filters | title | Exact / structured | Explicit (boolean / exact match) |
The number of combinations becomes a bit absurd when you start actually thinking about it. That’s before we even start talking about combining methods through fusion or reranking (that’s a future me problem :) )
UI/UX
The front-end is simplistic, both visually and in terms of structure. The landing page leads to 3 additional screens.
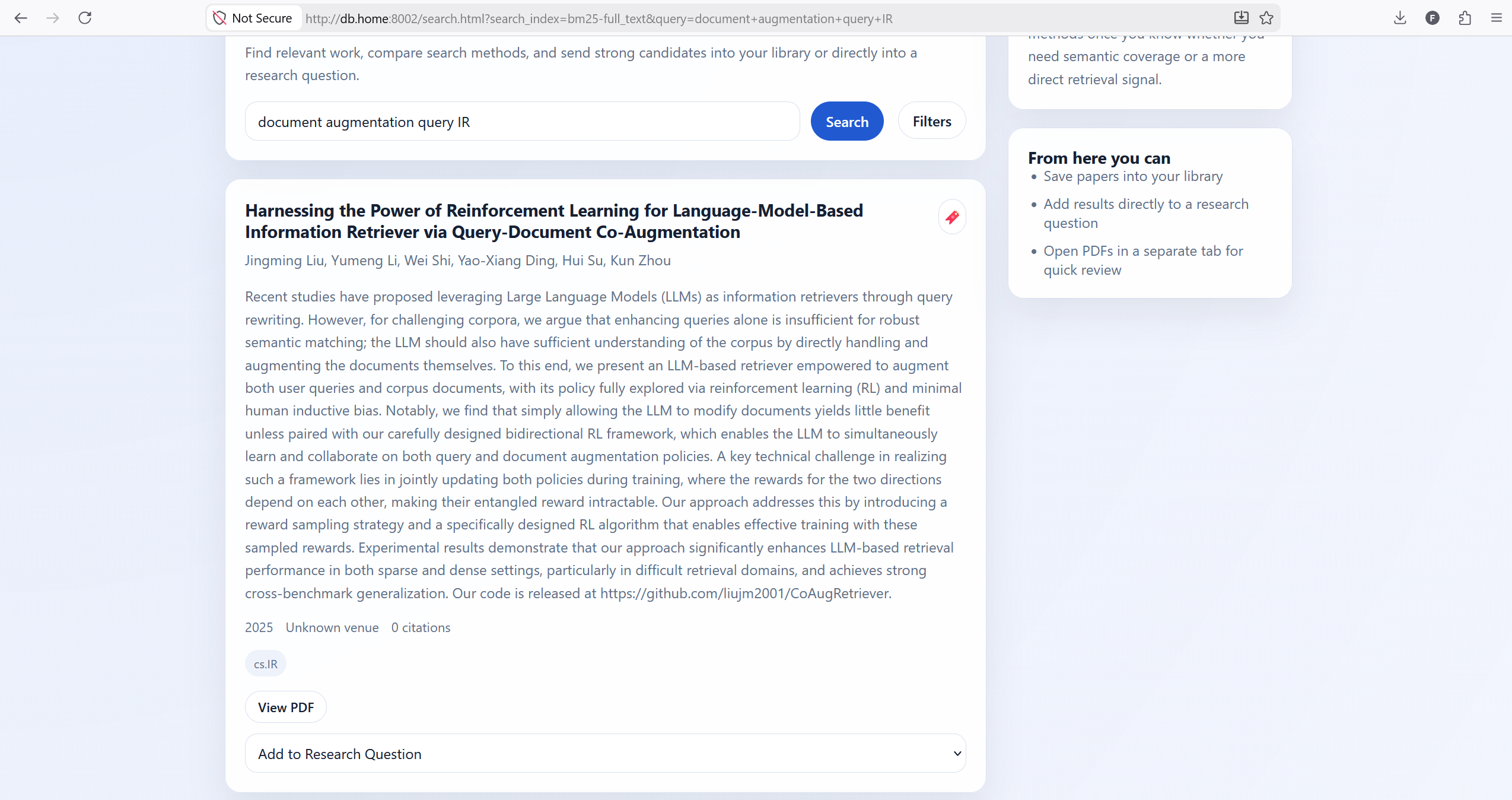
The search page is where the magic happens. Users can enter their queries up top. The results are presented in cards that have a red bookmark icon and a drop-down that allows them to add questions to a given RQ.
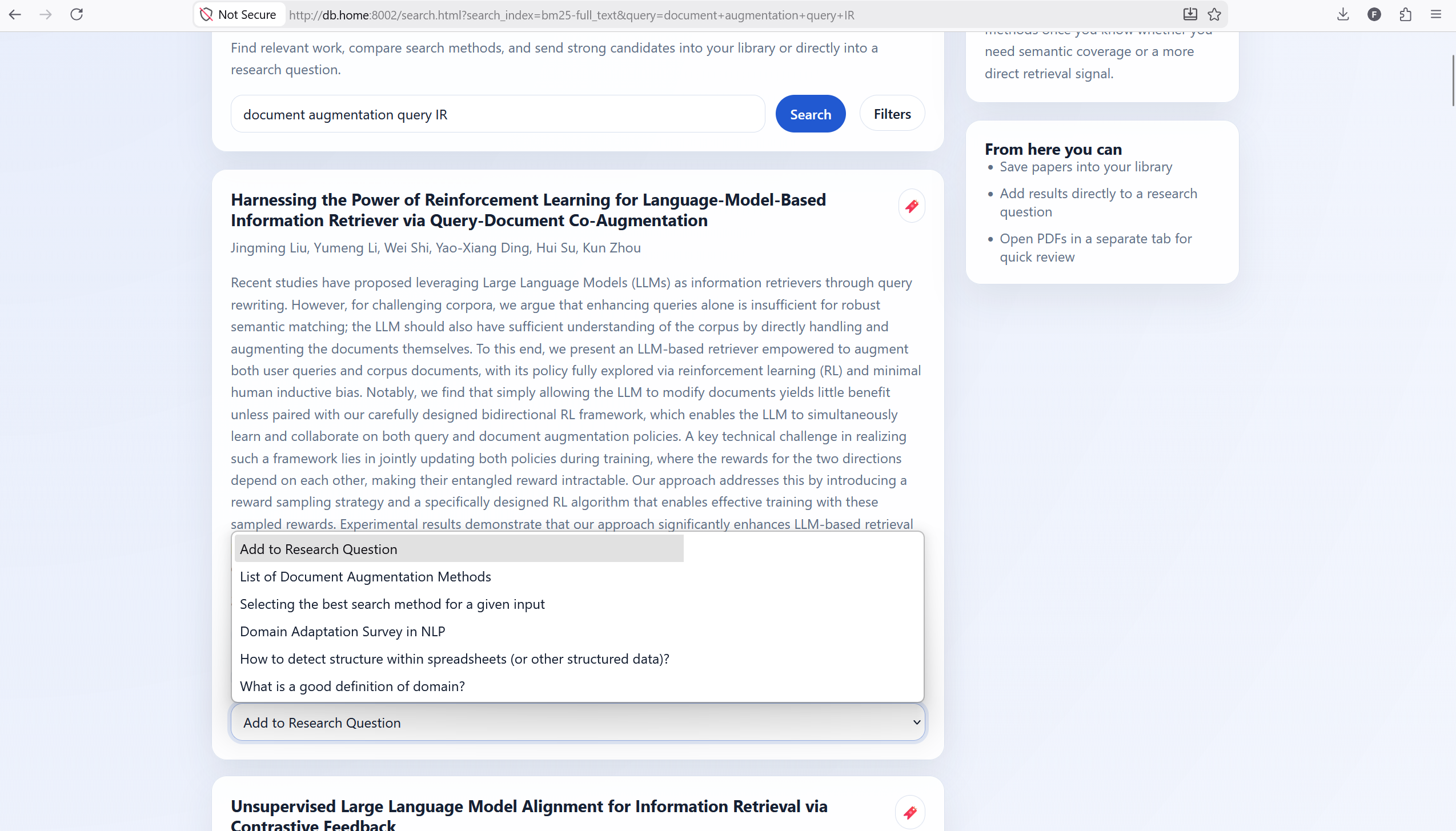
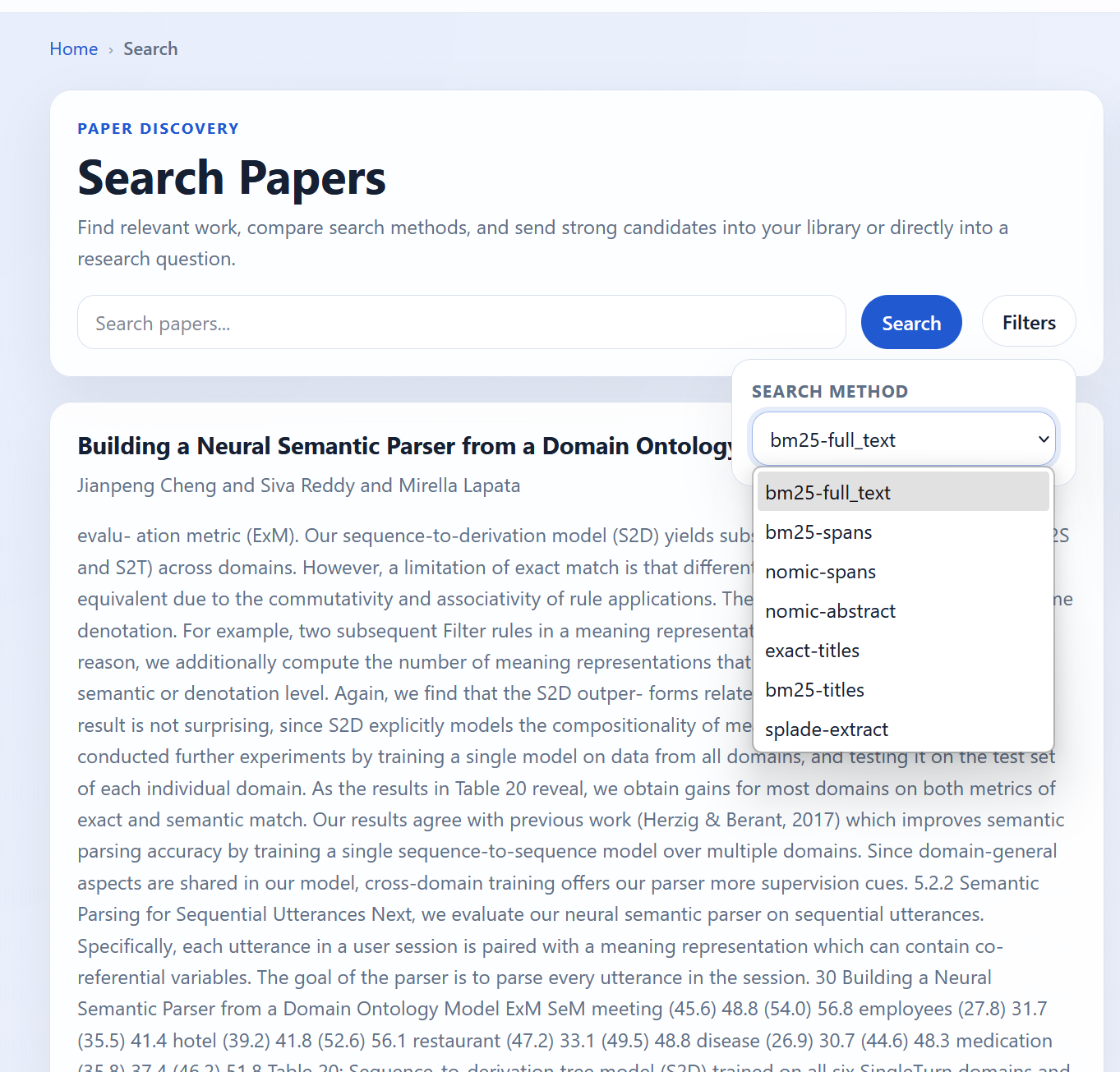
By clicking on filters, users have the ability to select a large number of index + source combinations. There are currently no other filters, but I plan on adding filters and sorts on citation count, year of publication, and other such fields.
Curation
Ok so you like a search result, what next? I created two areas to store results.
- A library (I assume you are familiar with the concept)
- A repository of research questions
Library
Library is still pretty rudimentary, but acts as a store of papers I want to read or have read.
It’s currently limited to adding papers, removing papers, or adding them to RQs. I’m currently using the library to triage papers until they find a more permanent home elsewhere. Eventually I’d like to have more filters and search over the library. This hasn’t really been a high priority since I’m primarily using the tool to organize research questions.
Research Question Management
Research questions is the term used, but this is a placeholder for any way of organizing results to any open questions I have. There is first a page that has the research questions themselves.
Upon clicking on a RQ, you can see the papers associated with that research question. There are also recommendations on the right-hand side.
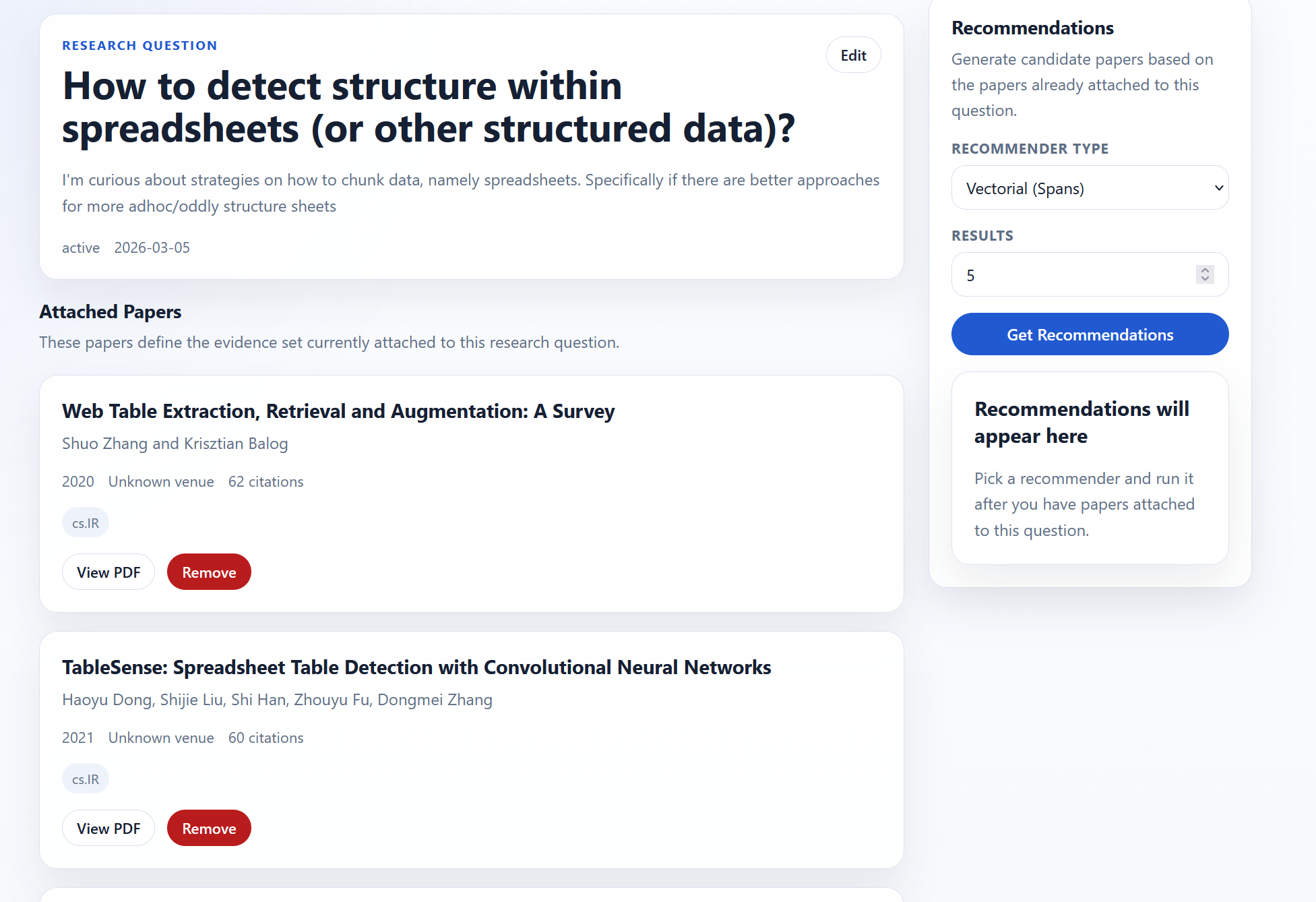
There are two recommendation engines:
- vector-based
- bibliographic-based
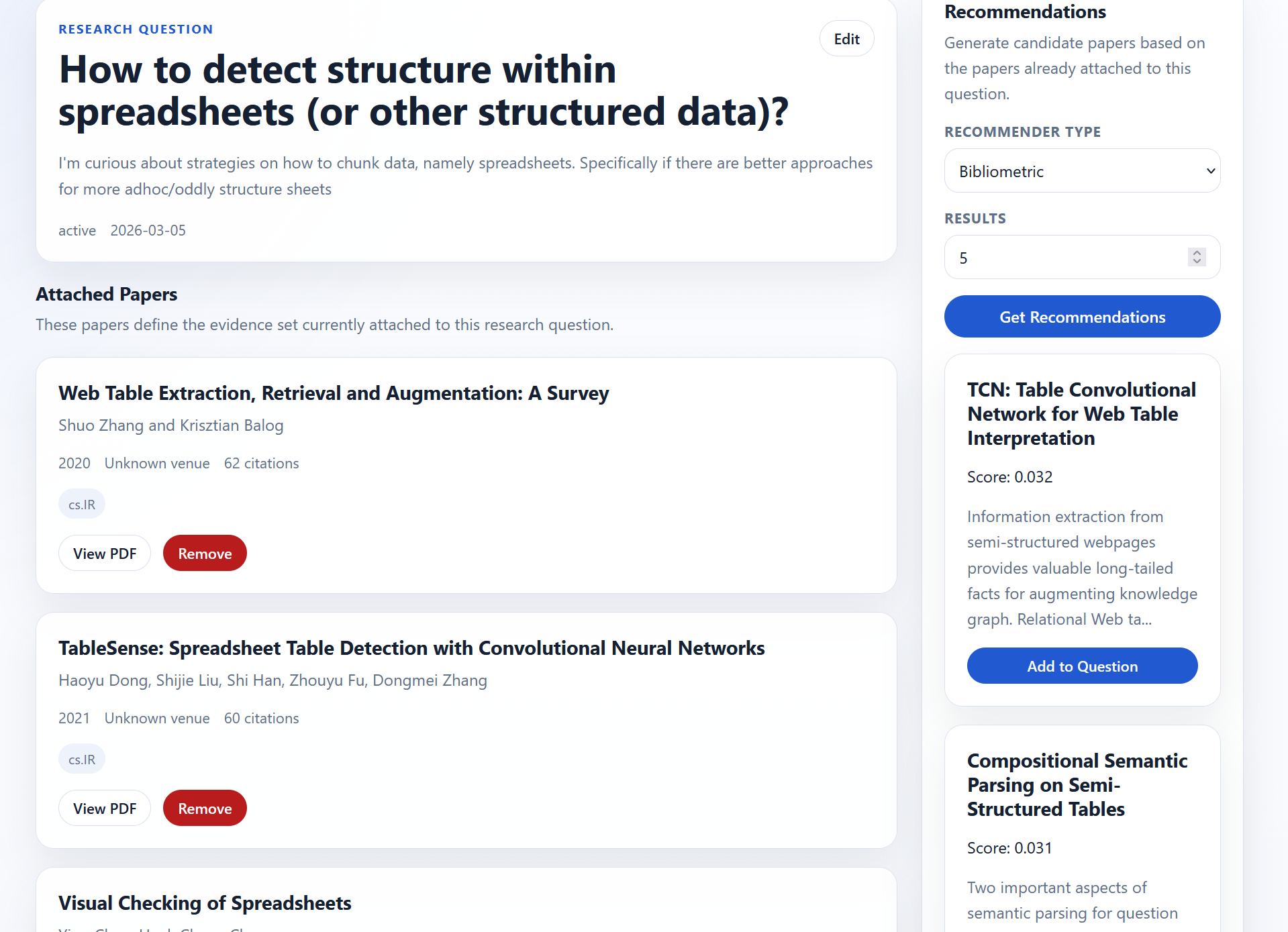
Under the hood the methods are pretty simple:
- Vector (Abstract)
- Pull in all papers for the RQ
- Aggregate the vectors for each paper (mean, max, etc.)
- Use that aggregate to find relevant papers
- Return top-k excluding the papers shared
- Bibliographic
- Run bibliographic similarity across all papers in the RQ
- Use RRF to fuse the rankings
- Return top-k excluding the papers shared
I eventually want to add more recommendation methods, but these two have worked well enough to start.
Next Steps
This tool was a good first stab, but there are still a number of issues that need to be addressed and potential improvements.
Large Scale Usage
I have used this tool for 5 RQs so far. It’s worked pretty well so far and surfaced papers I hadn’t found through the aforementioned tools. That being said, 5 RQs is not a lot and even the largest only has 9 papers as of writing. I’m currently planning a large-scale survey for definitions of domain in NLP which I expect to net 100+ papers so that will likely be a good test of how the system performs at scale. Currently there isn’t the ability to search and pagination was an afterthought. I expect the UI/UX to suck a bit due to that, but that is easily addressable.
Data Analysis and Data Gathering
I used a straightforward approach to create the search corpus and monitor its health. The biggest shortcomings I see in the intake pipeline are the following
- A single source was used (arXiv), so some papers or other works are missing. It’s also harder to create valid citations since the final venue isn’t published.
- The ingestion pipeline could use additional work
- Currently tags are used to filter relevant papers, but it’s a naive heuristic. I’m in the process of training a model to score relevance. A first attempt didn’t work reliably (a lot of false negatives), so I’m currently labeling sampled abstracts and fine-tuning a BERT-like model.
- The pipeline fails occasionally and there isn’t really a mechanism to identify failure points or recover from them. Note that the failure rate is tracked, but currently my system makes it hard to fix individual failures
- The semantic search API used to pull in citation and reference information seems to be missing some data as some papers’ arXiv IDs may not be pulled in. I want to replace their service with my own (one day)
- Structure isn’t used to pull in text or chunk it, so some of the boundaries for spans don’t make a lot of sense. I’m currently writing an LLM-based chunker
- There are no individual or corpus level quality checks
- At the individual level
- No checks for paper quality metrics such as methodological issues, misleading citations, etc.
- Some preprints are very wonky, there is everything from class projects to rants
- Spans are sometimes irrelevant text and there isn’t a tag or way to verify that
- Corpus level
- Dedupe, I don’t think this is an issue yet based on what I’ve seen this could easily become an issue if there are multiple sources
- What percentage of data is missing or not indexed isn’t currently tracked
- At the individual level
UI/UX
The underlying premise of this project is that it’s hard to choose the right search method. Part of the reason is that using the methods correctly isn’t intuitive. Keyword methods require choosing keywords relevant to your query and in your corpus. Neural methods are sensitive to word choice. Query augmentation approaches should be corpus aware and reflect what words are being added to the user. Each method has its quirks. How to interface with each method should be built into the UI. I think there are a couple ways of doing that
- Query phrasing suggestion methods
- For keyword methods suggest synonyms and/or often used words using corpus statistics
- For neural methods use an n-gram LLM over the fine-tuning corpus to propose good searches (making sure the input resembles the training data as much as possible)
- For query augmentation, integrating the approaches above and giving the user a way to understand if/how query augmentation is being used
- Use of an agent as a router to abstract method selection from the user. I propose a potential mechanism for this in the next section
More Search Tools
The approaches used to find relevant papers so far are limited in number and scope. Here are additional approaches I’d like to implement
- Cross-encoder (as a reranker)
- Listwise prompt-based reranker
- Pairwise prompt based reranker
- ColBERT and the subsequent works to reduce resource usage
- Additional dense embeddings, including custom dense embeddings that allow for different question formats (e.g., research questions -> spans)
- Query rewriting/augmentation, e.g., DocT5query or
- Document augmentation
- PRF and other “expansion” methods
Most other methods can be supported with my current stack, while others not so much. For example ColBERT requires additional indexing not available in paradedb. They just require additional code or training additional models. As we increase the number of tools, the number of pipelines increases exponentially and we need to handle this in a straightforward way.
DSL for Search
One of the things that’s hard about search is that it isn’t a single task, it’s a large myriad of tasks masquerading as one like similarity. A given query from different users could have different valid results, different inputs could map to different results. User intent is hard to decipher from just a query. Not all searches are as simple as just finding results, like tip of the tongue search, exploration, recommendation, and question answering for example. It seems that no single approach to search works best (in the context of RAG, but I think this result is universal). You’re not going to do QA and simple keyword retrieval with the same tool. For that reason intent needs to be understood and the user’s intent needs to be potentially clarified through direct interaction.
Current LLMs can write code (to a certain extent), so why not have code to create search pipelines optimized for each problem? A DSL (domain-specific language) for search and recommendations would be a good opportunity for this. It provides composability, inspectability and reproducibility.
Moreover a DSL can allow for a compact way of constructing and quickly trying out different pipelines without needing to start over from scratch. A lot of the components could be recombined in new and different ways. A DSL can allow for experimentation and optimization in a way that isn’t otherwise possible.
Beyond Search
Beyond search, there are other techniques to find relevant papers. One idea I’ve been toying with is that we can use workflows to do systematic surveys over the entire corpus.
The approach is simple
- define criteria for relevant papers that can be translated into tags/classes
- design a workflow that uses simple conditional logic to filter results
- implement the pipeline
- implement skeleton for pipeline
- create initial classification prompts
- run pipeline
- review each classification stage to make sure that results are adequate
- review the resulting papers and sample rejected papers to ensure the false positives were limited
This approach allows you to cast a wider net, estimate global performance (risk of non-inclusion) using traditional metrics. This likely isn’t a new approach, but I think the trick here is how to make this approach work reliably and quickly (i.e., not annotating every single paper). I’m currently trying this out for my survey approach and seeing if this is a viable approach.
Closing Remarks
Overall I’m happy with the preliminary results. A bit meta, but as I was researching and verifying claims for this post I used the tool a few times. For me this work solves the search and organization problems I was having with my former approach. I plan on sharing the underlying code when it’s further along. I haven’t productionalized the code and the front-end is entirely vibe coded. It’s got some jagged edges. I’m currently using the tool to survey definitions of domains in NLP, so there will be more updates (and potentially a code base) after I get done with that.